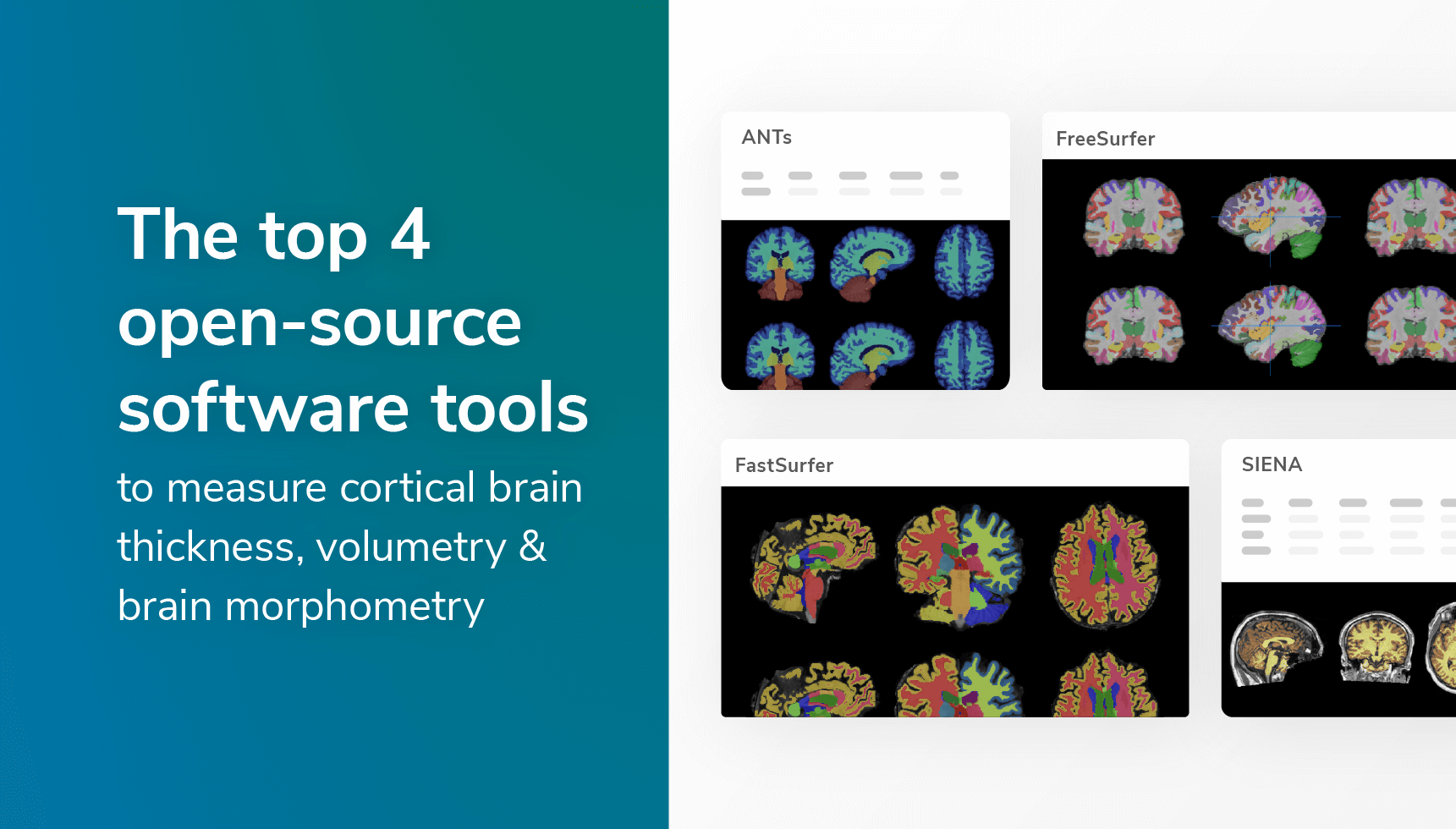
Imaging trends and tools are evolving more than ever. Many companies and technicians are developing new algorithms to help researchers and clinical professionals diagnose and measure disease progression efficiently.
Many tools are being developed every day, and even though some tools that we are going to discuss were made years ago, they are still the standard. This shows how complicated it is to improve current results, widely accepted by the scientific community. That is the case with the following open-source tools.
Powerful open-source tool for registration and normalization of brain imaging. Has a powerful tool for brain segmentation for six different tissues and cortical thickness computation. Can be used to apply cortical labeling using any anatomical atlas and the proper brain template. Studies show that it has a higher predictive performance than FreeSurfer (https://pubmed.ncbi.nlm.nih.gov/24879923/).
This is probably the most widely used open-source tool for structural, functional, and diffusion brain imaging. It is the standard tool for brain segmentation and volumetry computation. The software’s citations are already more than 6500 in scientific publications (source: https://www.sciencedirect.com/) . The segmentation pipeline also includes hippocampal subfields segmentation, brainstem substructures segmentation, longitudinal volumetric analysis, and more. This tool is also computationally expensive, so it might take a long time to process.
Built on top of the FreeSurfer brain reconstruction method, accelerates considerably the computation of brain morphometry. Open-source validated machine learning algorithm that can compute the bran morphometrics in a fraction of the time. It can mimic FreeSurfer’s brain segmentation and parcellation in less than one minute and cortical surface reconstruction in less than an hour.
Cross-sectional and longitudinal brain analysis compute brain volume from a single T1 image. It is also widely used. Part of the FSL library, which also contains many other imaging capabilities on a wide range of imaging modalities.
|
Tool
|
Primary Use
|
Computation Time
|
Key Strength
|
Platform Access
|
|
FreeSurfer
|
Segmentation, volumetry, cortical thickness
|
10–24 hours per subject
|
Gold standard; 6,500+ citations; widest adoption
|
✓ Available on QMENTA
|
|
ANTs
|
Registration, normalisation, cortical thickness
|
Moderate
|
Higher predictive performance than FreeSurfer in some tasks
|
✓ Available on QMENTA
|
|
FastSurfer
|
Segmentation, parcellation (FreeSurfer-compatible)
|
<1 min (segmentation); <1 hr (full surface)
|
Deep learning speed; ideal for large-scale studies
|
✓ Available on QMENTA
|
|
SIENA (FSL)
|
Brain volumetry, atrophy measurement
|
Moderate
|
Cross-sectional and longitudinal atrophy; part of FSL ecosystem
|
✓ Available on QMENTA
|
Why These Tools Are Hard to Use Alone — and How QMENTA Solves It
It can be quite challenging to use these kinds of tools by yourself. You have to read and understand the documentation, organize your data properly so the tool knows how to read it, write the command or build a script that will run the tool, and have the proper system configuration…
It can also cause a mess in your computer as the amount of files produced by these tools is not small. It can be hard to do a proper analysis and a pain to navigate through the myriad of results that each tool can produce. Managing the data adds a lot of burden in a task that should be simple. It is prone to error and resource-consuming.
QMENTA can help you with that because all the previous tools are available in the QMENTA platform. You can just plug your data into the platform and start analyzing. It takes seconds to start the analysis and just a few clicks. All the results are saved afterwards and organized in your project for you to inspect.
Frequently Asked Questions
What is the best open-source tool for cortical brain thickness measurement?
The two most commonly used open-source tools for cortical thickness measurement are FreeSurfer and ANTs (Advanced Normalization Tools). FreeSurfer is the most widely used and is cited in over 6,500 scientific publications, making it the de facto standard for brain segmentation, parcellation, and cortical thickness computation. ANTs is a powerful alternative for registration and normalisation, with studies showing it has higher predictive performance than FreeSurfer for cortical thickness in certain applications. FastSurfer, which runs on top of FreeSurfer's reconstruction method, offers a deep learning-accelerated alternative that produces equivalent results in significantly less computation time.
What is FreeSurfer and what can it measure?
FreeSurfer is an open-source software suite for processing and analysing human brain MRI data, developed at the Martinos Center for Biomedical Imaging at Massachusetts General Hospital. It is the most widely used tool for structural brain imaging and is cited in over 6,500 scientific publications. FreeSurfer can compute cortical thickness, surface area, cortical volume, subcortical volume, hippocampal subfield segmentation, brainstem substructure segmentation, and longitudinal volumetric analysis from T1-weighted MRI scans. It requires substantial computation time and technical configuration but produces highly validated, reproducible outputs.
What is FastSurfer and how does it differ from FreeSurfer?
FastSurfer is an open-source deep learning-based brain segmentation and cortical parcellation tool built on top of FreeSurfer's brain reconstruction methodology. It is designed to produce FreeSurfer-compatible results in a fraction of the computation time — brain segmentation and parcellation can be completed in under one minute, and full cortical surface reconstruction in under one hour, compared to 10–24 hours for standard FreeSurfer processing. FastSurfer uses a convolutional neural network (CNN) for whole-brain segmentation, making it particularly valuable for large-scale studies or workflows where processing speed is a constraint.
What is SIENA and what type of brain analysis does it perform?
SIENA (Structural Image Evaluation using Normalisation of Atrophy) is a brain volumetry tool within the FSL (FMRIB Software Library) suite. It performs cross-sectional brain volume analysis from a single T1-weighted MRI and longitudinal analysis by comparing two timepoint scans to measure atrophy rates. SIENA is widely used for measuring whole-brain atrophy in neurodegenerative disease studies, particularly in multiple sclerosis and Alzheimer's disease research. It is part of the broader FSL library, which also provides tools for functional MRI, diffusion MRI, and other modalities.
Why is it challenging to use these brain morphometry tools independently?
Running FreeSurfer, ANTs, FastSurfer, or SIENA independently requires reading extensive technical documentation, correctly organising input data in the expected formats, writing command-line scripts or pipelines, configuring the operating system environment, and managing large output file structures that can be difficult to navigate. These barriers mean that a significant portion of researcher time is spent on data management and technical configuration rather than scientific analysis. Each tool also produces different output formats, making it difficult to combine results across tools without custom integration work.
Can all four brain morphometry tools run on a single platform?
Yes — FreeSurfer, ANTs, FastSurfer, and SIENA (FSL) are all available on the QMENTA platform. Researchers can upload T1-weighted MRI data directly and run any of these tools through a point-and-click interface without installing software locally or writing command-line scripts. Results are automatically organised within the project, stored securely in the cloud, and accessible for downstream analysis and collaboration. This removes the technical setup burden and allows research teams to focus on analysis rather than infrastructure.