Jean-Martin Charcot first described ‘La sclérose en plaques disséminées’ and its associated neurologic triad - the combination of nystagmus, intention tremor and scanning speech - in 1868. Today, we know this condition to be Multiple Sclerosis (MS), a chronic inflammatory disease of the central nervous system affecting 2.5 million individuals worldwide. Histopathology reveals it is characterized by focal demyelinating lesions in the brain and spinal cord, with a clinical presentation that is dependent on the CNS regions affected.
The introduction of imaging in the second half of the 20th century enabled pathologic lesions to be detected in vivo. MRI in particular, with its superior contrast over CT, conceded greater sensitivity to the disease than clinical assessments alone. Quantitative and qualitative evaluation of MS was possible for the first time. MRI findings now form an essential component of the 2017 MAGNIMS/McDonald criteria for diagnosing MS and are key to tracking disease progression and responses to novel therapies.
The Challenges of White Matter Lesion Segmentation
Manual labeling of MRI images remains the most accurate method for MS white matter lesion segmentation (WMLS). But this process can considerably infringe upon a radiologist’s workload. Lesion loads vary from high to low, sizes from a few mm3 to cm3, and shapes from ovoid to irregular. Studies place the average time for the task to be performed on a single 3D scan as high as 50 minutes! [1]
What’s more, manual segmentation results are error-prone and susceptible to high inter- and intra-rater variability, intensified by differences in the scanner, acquisition sequence, and image quality. Such shortcomings have prompted the development of automated methods for MS WMLS, several of which are available in the QMENTA cloud platform and delineated below.
MS White Matter Lesion Segmentation in the QMENTA platform
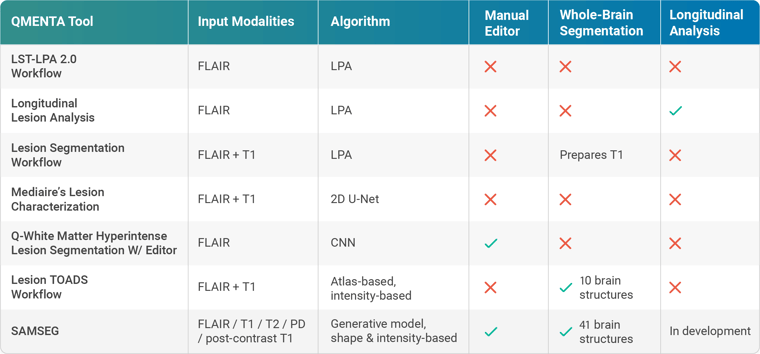
MRI-based MS lesion assessment typically utilizes hyperintensities in T2 FLAIR images. QMENTA’s LST-LPA 2.0 Workflow requires only a single FLAIR input to output estimated lesion maps, counts, and volumes. It implements SPM’s LST toolbox in the form of the lesion prediction algorithm (LPA) [2]. A large-scale logistic regression model alongside data from 53 cases of severe MS was used to train the algorithm. Its use has been validated across scanners and even in other diseases, such as Alzheimer’s. A longitudinal pipeline (Longitudinal Lesion Analysis) is also offered where different time points are available to facilitate assessments of lesion dissemination throughout both space and time.
LPA finds further utility in lesion-filling methods. Cortical atrophy is accelerated in MS patients and is shown to correlate with measures of clinical disability, but volume estimates can be biased in brains with high lesion loads. The Lesion Segmentation Workflow uses a lesion-filling approach to replace the lesion intensities derived via LPA performed on FLAIR and thus generates a T1 image that can be more accurately segmented thereafter.
Deep learning algorithms have also been recently suggested for WMLS, though with generally inferior results compared to human raters. In 2020 QMENTA partnered with mediaire to integrate their CE-marked Lesion Characterization module, among others. Although deep learning-based, they show that their algorithms are in fact on par with neuroradiologists in terms of segmentation accuracy [3]. Their 2D U-Net-like architecture is trained on axial, sagittal, and coronal FLAIR slices, subsequently employing a unanimous voting model which only accepts lesions that are present in all three orientations. Accepted lesions are then classified by region according to MAGNIMS/McDonald criteria to enable differentiation of MS lesions from age-related hyperintensities. The mediaire method ranked 1st in MICCAI 2021’s MSSEG-2 challenge [4].
We at QMENTA have developed our own deep learning-based tool [5], with the added feature of an in-app editor designed to further improve segmentation accuracy in FLAIR images: Q-White Matter Hyperintense Lesion Segmentation W/ Editor.
The prestige of supervised approaches is somewhat weakened by their poor generalizability. By nature, the relationship between images and labels in the training set is directly encoded into the algorithm, and for this reason, results may be limited when applied to images acquired with different scanners/protocols.
To side-step this caveat, the Lesion TOADS Workflow utilizes an unsupervised classifier [6]. Here lesions are modeled as an additional tissue class that shares its spatial distribution prior with healthy white matter. This takes place within the context of a topology-preserving atlas-based framework, which essentially enables simultaneous whole-brain and MS lesion segmentation of the combined T1-FLAIR input. It bypasses the two-phase approach necessary with lesion filling, making it one of the few dedicated tools for characterizing atrophy patterns in MS.
Finally, we introduce FreeSurfer’s SAMSEG [7] to the QMENTA platform. This tool completely decouples computational models of anatomy from models of the imaging process to accept a wide variety of hardware, protocols, and sequences, including pre-and post-contrast T1 images. Simultaneous whole-brain and MS WMLS are possible through the augmentation of its generative model with a binary lesion map. 41 brain structures are segmented (versus Lesion TOADS’ 10), with robust results even in controls. Further, MS lesion maps may be edited for false positives and negatives via a platform-based viewer.
References:
[1] Storelli, L., Pagani, E., Rocca, M. A., Horsfield, M. A., Gallo, A., Bisecco, A., Battaglini, M., De Stefano, N., Vrenken, H., Thomas, D. L., Mancini, L., Ropele, S., Enzinger, C., Preziosa, P., & Filippi, M. (2016). A Semiautomatic Method for Multiple Sclerosis Lesion Segmentation on Dual-Echo MR Imaging: Application in a Multicenter Context. AJNR. American Journal of Neuroradiology, 37(11), 2043–2049.
[2] Paul Schmidt. Bayesian inference for structured additive regression models for
large-scale problems with applications to medical imaging. PhD thesis, Ludwig Maximilians-Universität München, January 2017.
[3] Hitziger, S., Ling, W. X., Fritz, T., D'Albis, T., Lemke, A., & Grilo, J. (2022). Triplanar U-Net with lesion-wise voting for the segmentation of new lesions on longitudinal MRI studies. Frontiers in neuroscience, 16, 964250.
[4] https://portal.fli-iam.irisa.fr/msseg-2/
[5] Puch, Santi & Sanchez, Irina & Hernàndez, Aura & Piella, Gemma & Prčkovska, Vesna. (2019). Global Planar Convolutions for improved context aggregation in Brain Tumor Segmentation. arXiv:1908.10281
[6] Shiee, N., Bazin, P. L., Ozturk, A., Reich, D. S., Calabresi, P. A., & Pham, D. L. (2010). A topology-preserving approach to the segmentation of brain images with multiple sclerosis lesions. NeuroImage, 49(2), 1524–1535.
[7] Cerri, S., Puonti, O., Meier, D. S., Wuerfel, J., Mühlau, M., Siebner, H. R., & Van Leemput, K. (2021). A contrast-adaptive method for simultaneous whole-brain and lesion segmentation in multiple sclerosis. NeuroImage, 225, 117471.

Frequently Asked Questions
What is automated white matter lesion segmentation and why does it matter for MS?
White matter lesion segmentation is the process of identifying and delineating areas of white matter hyperintensity in MRI images — bright regions visible on T2-FLAIR sequences that are characteristic of MS demyelination. Manual segmentation by a radiologist remains the most accurate method but is time-consuming, with studies placing average manual segmentation time at up to 50 minutes per 3D scan. It is also susceptible to inter- and intra-rater variability due to differences in scanner hardware, acquisition sequences, and image quality. Automated methods address these limitations by providing faster, more reproducible outputs at scale.
What is the LST-LPA algorithm and what does it require as input?
The Lesion Prediction Algorithm (LPA), implemented through the SPM-based LST toolbox, is one of the most widely validated automated MS lesion segmentation tools. It requires only a single FLAIR input to generate estimated lesion maps, lesion counts, and lesion volumes. The algorithm was trained using logistic regression on data from 53 severe MS cases and has since been validated across different scanner types and even applied to other conditions such as Alzheimer's disease. A longitudinal pipeline variant enables assessment of lesion changes across multiple timepoints.
How do deep learning approaches to MS lesion segmentation compare to traditional methods?
Deep learning-based lesion segmentation algorithms have demonstrated accuracy approaching that of expert neuroradiologists in some studies. The mediaire Lesion Characterization module — integrated into QMENTA and CE-marked — uses a 2D U-Net architecture trained on axial, sagittal, and coronal FLAIR slices with a unanimous voting model, and ranked first in the MICCAI 2021 MSSEG-2 challenge. Deep learning approaches generally show superior speed and can match neuroradiologist accuracy, but their performance can be limited when applied to images from scanners or protocols not represented in their training data — a limitation known as poor generalisation.
What is SAMSEG and how does it handle multi-sequence MRI for MS?
SAMSEG (Sequence Adaptive Multimodal SEGmentation), developed within FreeSurfer, is a generative probabilistic segmentation tool that decouples its computational models of brain anatomy from the imaging acquisition process. This makes it tolerant of a wide variety of MRI hardware, protocols, and sequences — including pre- and post-contrast T1 images — without requiring retraining. It segments 41 brain structures simultaneously with whole-brain and MS lesion segmentation, and allows MS lesion maps to be edited for false positives and negatives through a platform-based viewer. Its results in healthy controls are robust, which is valuable for studies requiring comparison to normative populations.
What is the Lesion TOADS workflow and why is it suited for measuring atrophy in MS?
Lesion TOADS uses an unsupervised classifier that models MS lesions as an additional tissue class sharing its spatial distribution with healthy white matter, within a topology-preserving atlas-based segmentation framework. This approach allows simultaneous whole-brain segmentation and MS lesion segmentation from combined T1 and FLAIR inputs, bypassing the two-phase lesion-filling approach required by other tools. Because it directly addresses lesion-related bias in volumetric measurements, it is one of the few dedicated tools for accurately characterising cortical atrophy patterns in MS — a clinically important measure that correlates with disability progression.
